A team of researchers has generated a developmental map of a key sound-sensing structure in the mouse inner ear. Scientists at the National Institute on Deafness and Other Communication Disorders (NIDCD), part of the National Institutes of Health, and their collaborators analyzed data from 30,000 cells from mouse cochlea, the snail-shaped structure of the inner ear. The results provide insights into the genetic programs that drive the formation of cells important for detecting sounds. The study also sheds light specifically on the underlying cause of hearing loss linked to Ehlers-Danlos syndrome and Loeys-Dietz syndrome.
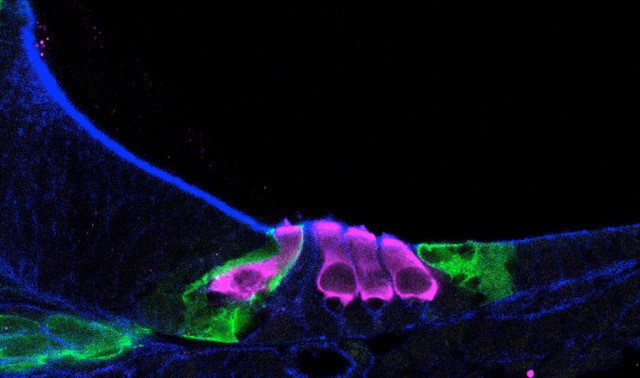
Single-cell RNA sequencing helped scientists map how sensory hair cells (pink) develop in a newborn mouse cochlea.
Source: Helen Maunsell, NIDCD/NIH
The study data is shared on a unique platform open to any researcher, creating an unprecedented resource that could catalyze future research on hearing loss. Led by Matthew W. Kelley, Ph.D., chief of the Section on Developmental Neuroscience at the NIDCD, the study appeared online in Nature Communications. The research team includes investigators at the University of Maryland School of Medicine, Baltimore; Decibel Therapeutics, Boston; and King’s College London.
“Unlike many other types of cells in the body, the sensory cells that enable us to hear do not have the capacity to regenerate when they become damaged or diseased,” said NIDCD Director Debara L. Tucci, M.D., who is also an otolaryngology-head and neck surgeon. “By clarifying our understanding of how these cells are formed in the developing inner ear, this work is an important asset for scientists working on stem cell-based therapeutics that may treat or reverse some forms of inner ear hearing loss.”
In mammals, the primary transducers of sound are hair cells, which are spread across a thin ribbon of tissue (the organ of Corti) that runs the length of the coiled cochlea. There are two kinds of hair cells, inner hair cells and outer hair cells, and they are structurally and functionally sustained by several types of supporting cells. During development, a pool of nearly identical progenitor cells gives rise to these different cell types, but the factors that guide the transformation of progenitors into hair cells are not fully understood.
To learn more about how the cochlea forms, Kelley’s team took advantage of a method called single-cell RNA sequencing. This powerful technique enables researchers to analyze the gene activity patterns of single cells. Scientists can learn a lot about a cell from its pattern of active genes because genes encode proteins, which define a cell’s function. Cells’ gene activity patterns change during development or in response to the environment.
“There are only a few thousand hair cells in the cochlea, and they are arrayed close together in a complex mosaic, an arrangement that makes the cells hard to isolate and characterize,” said Kelley. “Single-cell RNA sequencing has provided us with a valuable tool to track individual cells’ behaviors as they take their places in the intricate structure of the developing cochlea.”
Building on their earlier work on 301 cells, Kelley’s team set out to examine the gene activity profiles of 30,000 cells from mouse cochleae collected at four time points, beginning with the 14th day of embryonic development and ending with the seventh postnatal day. Collectively, the data represents a vast catalog of information that researchers can use to explore cochlear development and to study the genes that underlie inherited forms of hearing impairment.
Kelley’s team focused on one such gene, Tgfβr1, which has been linked to two conditions associated with hearing loss, Ehlers-Danlos syndrome and Loeys-Dietz syndrome. The data showed that Tgfbr1 is active in outer hair cell precursors as early as the 14th day of embryonic development, suggesting that the gene is important for initiating the formation of these cells.
To explore Tgfβr1’s role, the researchers blocked the Tgfbr1 protein’s activity in cochleae from 14.5-day-old mouse embryos. When they examined the cochleae five days later, they saw fewer outer hair cells compared to the embryonic mouse cochleae that had not been treated with the Tgfbr1blocker. This finding suggests that hearing loss in people with Tgfβr1 mutations could stem from impaired outer hair cell formation during development.
The study revealed additional insights into the early stages of cochlear development. The developmental pathways of inner and outer hair cells diverge early on; researchers observed distinct gene activity patterns at the earliest time point in the study, the 14th day of embryonic development. This suggests that the precursors from which these cells derive are not as uniform as previously believed. Additional research on cells collected at earlier stages is needed to characterize the initial steps in the formation of hair cells.
In the future, scientists may be able to use the data to steer stem cells toward the hair cell lineage, helping to produce the specialized cells they need to test cell replacement approaches for reversing some forms of hearing loss. The study’s results also represent a valuable resource for research on the hearing mechanism and how it goes awry in congenital forms of hearing loss.
The authors have made their data available through the gEAR portal (gene Expression Analysis Resource), a web-based platform for sharing, visualizing, and analyzing large multiomic datasets. The portal is maintained by Ronna Hertzano, M.D., Ph.D., and her team in the Department of Otorhinolaryngology and the Institute for Genome Sciences (IGS) at the University of Maryland School of Medicine.
“Single-cell RNA sequencing data are highly complex and typically require significant skill to access,” said Hertzano. “By disseminating this study data via the gEAR, we are creating an ‘encyclopedia’ of the genes expressed in the developing inner ear, transforming the knowledge base of our field and making this robust information open and understandable to biologists and other researchers.”
This press release describes a basic research finding. Basic research increases our understanding of human behavior and biology, which is foundational to advancing new and better ways to prevent, diagnose, and treat disease. Science is an unpredictable and incremental process; each research advance builds on past discoveries, often in unexpected ways. Most clinical advances would not be possible without the knowledge gained through basic research.
This research was supported by the NIDCD Division of Intramural Research (ZIADC000039) as well as (ZICDC000086) to the Genomics and Computational Biology Core, which is led by Robert Morell, Ph.D., and by King’s College London. The gEAR portal is primarily supported by the Hearing Restoration Project of Hearing Health Foundation, New York City, with additional funding from the NIDCD (R01DC013817) and from NIH’s National Institute of Mental Health (R24MH114815). Computational resources of the NIH HPC Biowulf cluster were used in this study.
About the National Institute on Deafness and Other Communication Disorders (NIDCD): The NIDCD supports and conducts research and research training on the normal and disordered processes of hearing, balance, taste, smell, voice, speech, and language and provides health information, based upon scientific discovery, to the public.
About the National Institutes of Health (NIH): NIH, the nation's medical research agency, includes 27 institutes and centers and is a component of the U.S. Department of Health and Human Services. NIH is the primary federal agency conducting and supporting basic, clinical, and translational medical research, and is investigating the causes, treatments, and cures for both common and rare diseases.

